More Information
Submitted: 23 September 2020 | Approved: 12 October 2020 | Published: 14 October 2020
How to cite this article: Lakshamanan S, Mahalakshmi S, Ramya H, Nidhi S. Effect of sperm DNA fragmentation on ICSI outcome: A prospective study. Clin J Obstet Gynecol. 2020; 3: 127-131.
DOI: 10.29328/journal.cjog.1001065
ORCiD: orcid.org/0000-0003-1468-8996
Copyright License: © 2020 Lakshamanan S, et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Keywords: Sperm; DNA; Embryo; Assisted reproductive technology; Blastocyst; ICSI
Effect of sperm DNA fragmentation on ICSI outcome: A prospective study
Saravanan Lakshamanan1, Saravanan Mahalakshmi1, Harish Ramya1 and Sharma Nidhi2*
1ARC International Fertility Center, 947, Avinashi Road, Puliakulam, Coimbatore-641037, Tamil Nadu, India
2Department of Obstetrics and Gynecology, Saveetha University,Kuthambaakam, Chennai-600124, Tamil Nadu, India
*Address for Correspondence: Sharma Nidhi, ARC International Fertility Center, 947, Avinashi Road, Puliakulam, Coimbatore-641037, Tamil Nadu, India, Tel: +91 9445560392; Email: [email protected], [email protected]
Aim and objectives: The primary aim was to measure the sperm DNA damage and to study the magnitude of sperm DNA damage. Secondary objective was to study the effect of sperm DNA fragmentation on Day 5 Blastocyst expansion (graded 1-5).
Results: There is an increase in sperm DNA fragmentation with an increase in age. Increased sperm DNA fragmentation is also associated with abnormal motility and morphology in semen samples. However, there is no reduction in expansion or grade of blastocyst.
Conclusion: Sperm DNA fragmentation testing is a useful investigation in unexplained infertility. However, Sperm DNA fragmentation has no significant association with Day 5 embryo grade in ICSI cycles.
Thesis work of Fellowship in Reproductive Medicine student: Dr. Ramya Harish
Semen analysis, traditionally done as per WHO 2010 includes sperm volume, liquefaction time, semen pH, sperm concentration, sperm morphology, semen fructose, sperm motility and semen reactive oxidative species (ROS). The WHO 1999 guidelines had also analyzed sperm agglutination tests and leukocyte concentration. These laboratory tests elaborates the morphology and function of human sperm, however this analysis may not reveal defects in sperm DNA [1-3]. It is interesting to note that only 15% of infertile men have normal semen analysis as per WHO guidelines [4]. Whether sperm DNA damage adversely affects reproductive outcomes in natural and assisted conception is a matter of debate. A few studies have shown that immature nucleoprotein and sperm DNA single and double strand breaks can lead to implantation failures, lower clinical pregnancy rates and early miscarriages [4]. More recently, several techniques designed to improve sperm selection for conventional ICSI have demonstrated to increase fertilization rates, enhance embryo quality after successful fertilization, and optimize pregnancy rates. DNA damage may occur in the form of single or double strand breaks, and both types can be analyzed and or quantified through different methods. These are the Sperm Chromatin Dispersion Test (SCDT), Sperm Chromatin Structure Assay (SCSA), deoxynucleotidyl transfer mediated dUTP nick end labeling assay (TUNEL) and single cell gel electrophoresis assay (COMET) [5,6]. Spermatogonia develop into primary spermatids and then into secondary spermatids, which shed their cytoplasm to form spermatozoon. DNA in spermatogonia is mainly histone bound and protamination of DNA takes place in testicular seminiferous tubules. During the process of sperm DNA maturation histones are replaced by protamine that are half the size of histones. In the mature spermatozoon nucleoprotein, 85% of mature sperm DNA is bound to protamine I and II and 15% of sperm DNA is bound to histones [7,8]. If histones are bound to > 15% of DNA it is termed as immature nucleoprotein and it has been postulated by a few studies as the most likely etiology of sperm DNA defects. The other etiological factors are varicocele, increased testicular temperature, stress, aging, increased BMI (Body Mass Index), smoking, and infrequent ejaculation [9,10]. Other factors are environmental pollution, male genital tract infections, toxins, radiation exposure, diabetes, testicular cancers and abuse of recreational drugs and caffeine. The biological mechanisms involved are defective protamination, increased apoptosis and oxidative stress. Defective protamination happens in the testes. Oxidative stress mostly happens as a result of sperm motility and metabolism while transit through the epididymis [11,12]. Apoptotic stress can happen either in testes or in the epididymis. Presence of leukocytes and immature sperm also add to the oxidative stress. Oxidative stress is sometimes also due to low antioxidants in the seminal plasma[13,14].
Sperm oxidative species are required in moderation for sperm capacitation, acrosome reaction, hyper activation and oocyte fusion however a high amount may produce sperm DNA damage. Sperm DNA fragmentation test may be indicated in work up of unexplained infertility, recurrent implantation failures, recurrent miscarriages, age > 40, varicocele and metabolic arrest of embryos at compaction stage. In human embryo the genome is oocyte dependent till day 3 after which the embryonic genome is activated. The activation of embryonic genome is a complicated process. The failure of embryonic genome to activate can lead to metabolic block of embryos on day 3. Hence, the primary objective of the present study was to identify the role of sperm DNA fragmentation testing in subfertile couples and correlate it with other traditional sperm analysis parameters. The secondary objective was to correlate sperm DNA fragmentation with embryo trophoectoderm and inner cell mass characteristics.
This is a prospective observational study conducted in the Department of Obstetrics and Gynecology of Saveetha medical college and ARC infertility hospital. The research proposal was approved and all collected data was examined and approved by the Institution Ethics Committee and review board. Informed written consent in local language was taken from all participant couples. The aim of the study was to investigate the effect of sperm DNA fragmentation in ICSI. Sixty couples (primary or secondary infertility) from period of one year were included in the study.
Male partners who had a sperm concentration of at least 1 x106/ml in raw semen were included. Couples with known gynecological pathology (e.g., known endometriosis, fibroids, and any previous operation to gynecological organs) were excluded. Fresh embryo transfer cycles and donor oocytes recipients were also excluded from the study.
Informed written consent was taken from Male partners who were attending infertility clinic for sperm DNA fragmentation testing by SCD (Sperm Chromatin Dispersion) method. Semen samples were collected by masturbation after 2–5 days of sexual abstinence. The semen was allowed to undergo liquefaction. DNA fragmentation was studied by SCD method (halo view, DNA fragmentation kit). Before testing, the sperm concentration is diluted to 5-10 million per ml (diluted by Vitromed V-sperm wash containing HEPES ,HTF AND HSF). If the sperm concentration was below 5million/ml the semen sample was centrifuged and the pellet was used for chromatin dispersion test.
The sperm chromatin dispersion test protocol has 5 steps
Step 1: Reagent preparation: In the first step all reagents are brought to the room temperature to prepare a working solution of Giemsa from the concentrated stock solution. The Giemsa working solution is made adding Giemsa stock solution with phosphate buffered saline and distilled water (1:1:1).
Step 2: Sample preparation: The agarose gel is melted at 90 degrees for 5 minutes in water baths. The agarose gel is now placed in 37 degrees water bath and allowed to equilibrate. Semen sample is added to fused agar. Immediately the semen agarose mixture is pipetted out smear is made on the slide and coverslip is placed. Now the slide is placed at 4 degrees for 5 min to produce a micro gel embedded with sperm cells.
Step 3: Denaturation and lysis: In this step the cover slip is gently removed and the sperm agarose smear is overlaid with acid denaturation solution for 7 minutes. After this the sperm agarose smear is overlaid with Lyses buffer for 25 minutes. Now the slide is washed with abundant distilled water for 5 minutes.
Step 4: Fixing and staining: Now the smear is dehydrated gradually by serially addition of 70% alcohol, followed by 90% alcohol and then followed by 100% alcohol for 2 minutes each and then air-dried. After fixing the Giemsa working solution prepared earlier is overlaid for 5-10 minutes to stain the smear. Strong staining is preferred to achieve easy visualization of the periphery of the dispersed DNA loop halos.
Step 5: Observation and Interpretation: The slide is observed under 40X bright field illumination. A minimum of 200 sperms are counted and scored on the pattern established by Fernandez, et al. Sperm DNA fragmentation (DFI%) is calculated as DNA fragmented spermatozoa/Total sperm counted. Table 1 gives the interpretation of halo size and sperm DNA fragmentation. Semen samples were prepared by density gradient method for ICSI. Magnetic assisted cell sorting was done when DNA fragmentation was more than 30%. Conventional ICSI was done and embryos were cultured till Day 5 with using Vitrolife single step media.
| Table 1: Interpretation of sperm DNA based on the size of halo. | |
| Size of the Halo | Interpretation |
| Big Halo | Normal |
| Medium Halo | Normal |
| Small Halo | Fragmented |
| No Halo | Fragmented |
| Degraded | Fragmented |
Age, motility and morphology, sperm DNA fragmentation and day 5-embryo grade (graded 1-5 based on the expansion of blastocyst, the inner cell mass and trophoectoderm morphology) was recorded.
Statistical analysis
Data was analyzed and graphical recording was done. Descriptive and inferential statistics were used to analyze the data. Comparisons of age, sperm count, sperm motility and sperm morphology was performed with the use of Karl Pearson Correlation Coefficient (r) was used for measuring the linear dependence of two variables. Correlation coefficient, -1 <= r <= 1, 1 represents strongly positively correlated, -1 represents strongly negatively correlated, 0 represents no correlation. A scatter plot was plotted using Cartesian coordinates to show values for two variables. Chi square test with Yate’s correction was used for analyzing categorical variables. Statistical analysis was done using the statistical software by MEDCALC (Belgium).
Mean age of participants was 35.83 + 4.81. It was found that with increased age the sperm count was not affected though motility and morphology were affected. As age increased there was increased sperm DNA fragmentation. A nonsignificant positive correlation was observed between DNA fragmentation and sperm motility. A nonsignificant negative correlation was also observed between fragmentation and morphology. Table 2 summarizes the correlation between demographic variables and semen parameters. Sperm DNA fragmentation also showed no effect on blastocyst expansion (Table 3). Increased sperm DNA fragmentation more than 30% was also not associated with trophoectoderm or inner cell mass grade (Tables 4,5).
| Table 2: Correlation of demographic characteristics and semen parameters {Pearson Correlation Coefficient (r) is used for measuring the linear dependence of two variables}. | |||||
| Sno. | Characteristic (x axis v/s y axis) | Coefficient of correlation(r) | Confidence interval | p value | Inference |
| 1 | Age v/s Sperm DNA fragmentation | +0. 3146 | 0.07-0.53 | 0.01436 | Positive significant |
| 2 | Sperm Count million/ml v/s Sperm DNA fragmentation | +0.03 | -0.23-0.28 | 0.8200 | Positive Non significant |
| 3 | Sperm Motility v/s Sperm DNA fragmentation | +0.0198 | -0.24-0.27 | 0.88064 | Positive Non significant |
| 4 | Sperm Morphology v/s Sperm DNA fragmentation | -0.0765. | -0.32-0.18 | 0.56384 | Negative Non significant |
| Table 3: Sperm DNA fragmentation (%) and Blastocyst Expansion Grade (p value=1, nonsignificant). | |||
| Blastocyst expansion | Grade 4 or 5 | Grade 3,2 or 1 | Total |
| SDF < 30% | 2 | 3 | 5 |
| SDF > 30% | 19 | 36 | 55 |
| Total | 21 | 39 | 60 |
| Table 4: Sperm DNA fragmentation and Inner cell mass Grade (p value = 0.3892, nonsignificant). | |||
| Inner cell Mass grade | Grade A | Grade B | Total |
| SDF < 30% | 4 | 1 | 5 |
| SDF > 30% | 34 | 21 | 55 |
| Total | 38 | 22 | 60 |
| Table 5: Sperm DNA fragmentation and Trophoectoderm Grade (p value=0.6586, nonsignificant). | |||
| Trophoectoderm Grade | Grade A | Grade B | Total |
| SDF < 30% | 2 | 3 | 5 |
| SDF > 30% | 23 | 32 | 55 |
| Total | 25 | 35 | 60 |
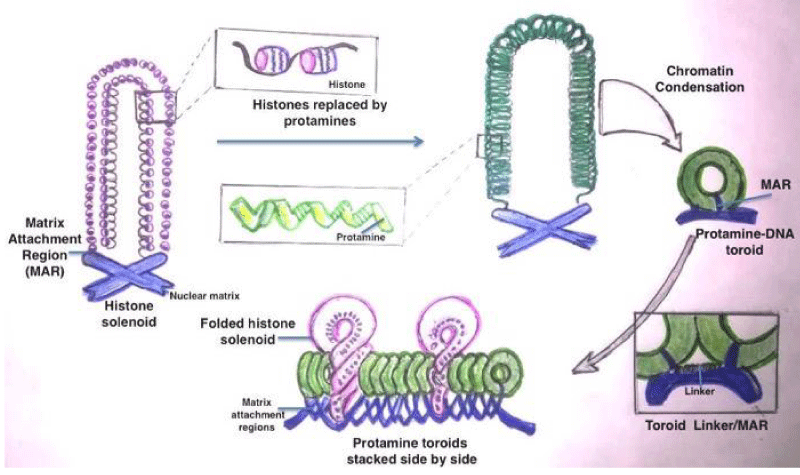
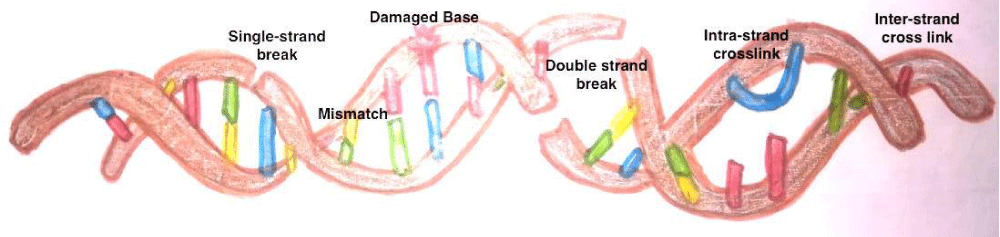
The sperm DNA contains Transition Nuclear proteins (TNPs) and protamine in toroids as apposed to histones in somatic cells and oocyte [15-17]. Figure 1 elaborates that the sperm DNA chromatin arrangement and replacement of histones with protamine during nucleoprotein maturation. The commonest defects are single stranded and double stranded DNA breaks. Other defects are base deletion and base modification. Besides there can be inter and intrastrand cross links (Figure 2).
Figure 1: Maturation of nucleoprotein and replacement of histones with protamine in Sperm Chromatins.
Figure 2: Types of Sperm DNA defects.
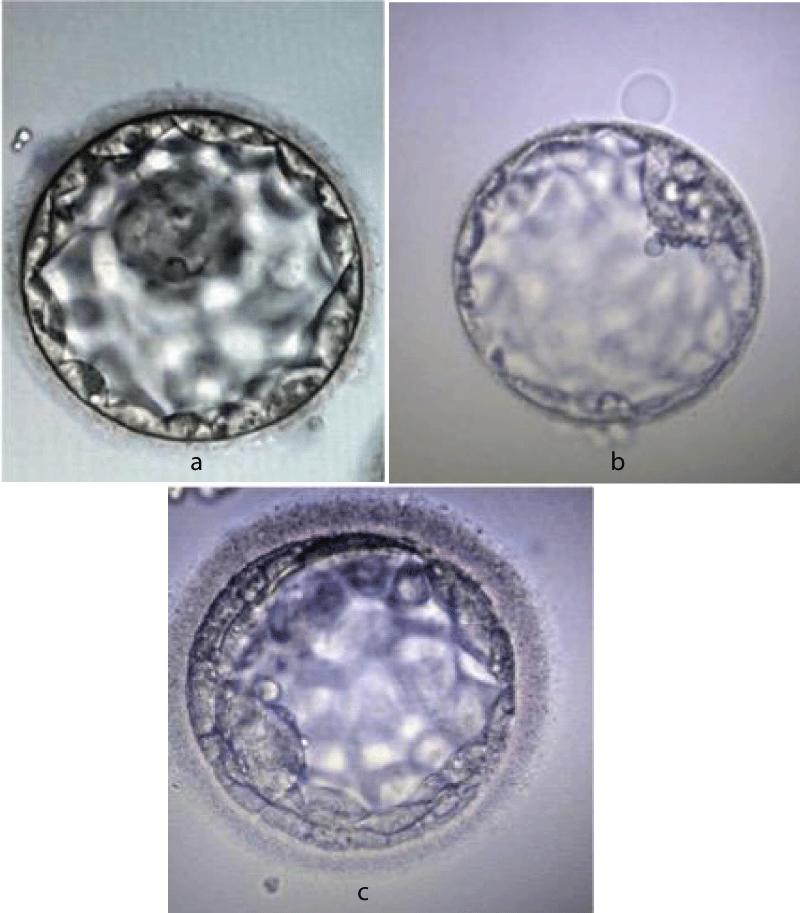
The protamination process has several advantages (a) DNA condensation results in a lighter nucleus to easy sperm ascent into the female genital tract (b) enhanced DNA stabilization from free radicals generated in seminal plasma as a result of sperm motility and metabolism (c) the somatic epigenetic genes are removed from the sperm nucleus enabling free reprogramming by the oocyte following syngamy (d) a check point in spermiogenesis (defects in protamination can act as a check point for onset of apoptotic pathways) (e) activation of oocyte following fertilization [18,19]. The embryos grading system (Gardener and Schoolcraft) is elaborated in figure 3a-c.
Figure 3: a: Blastocyst grade AA. b: Blastocyst grade BB. c: Blastocyst grade CC.
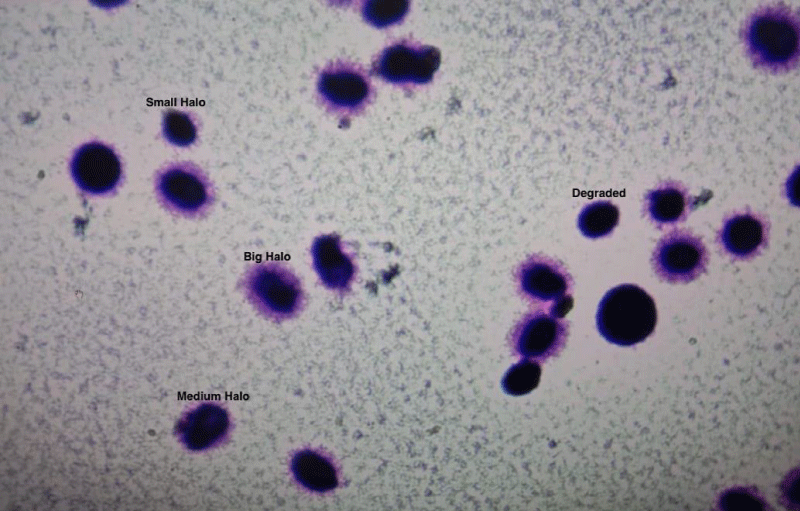
The process of protamination may at times be associated with damaged DNA. DNA fragmentation is a hallmark of apoptosis in human sperm, and may be caused by abnormal endonuclease activity and mediated by the plasma membrane protein Fas. Previous studies have demonstrated that the apoptosis is increased in the semen of infertile males. Apoptosis is also increased in samples with abnormal sperm morphology [10]. In accord with other studies, this study also suggests that with increased age, there is an increase in sperm DNA fragmentation. Figure 4 shows the spermatozoa halo test after staining.
Figure 4: Halo test for Sperm DNA fragmentation by Sperm Chromatin Dispersion test.
The oocyte has some repair mechanisms to repair the cracks in sperm DNA as evidenced by recent studies in rainbow trout oocytes and embryos [21,22]. Our study has also found that with an increase in sperm DNA fragmentation > 30% there is no effect on blastocyst expansion in human embryos. Inner cell mass and trophoectoderm grade also are not related to sperm DNA fragmentation (> 30%).
The use of sperm selection methods, for example density gradient centrifugation and swim-up, promote spermatozoa with the largest TEL-DNA (telomeric DNA) repeats to be selected that is helpful for improving embryo quality. Additionally, newer sperm selection methods like Magnetic assisted cell sorting used in high DFI samples can also improve ICSI outcome as evidenced by various studies in [23-25].
The commonest defects are single stranded and double stranded DNA breaks. Other defects are base deletion and base modification. Besides there can be inter and intrastrand cross-links.
Single strand breaks and abasic DNAs in the mature spermatozoa can be repaired by the oocyte with the help of BER (Base excision Repair) route before fusion of chromatids [26,27]. Double strand breaks can be repaired in the zygote by NHEJ (Non Homologous end joining) and HR (Homologous Recombination) pathways [28]. In the zygotic stage, the Non Homologous End Joining mechanism helps in the repair of double strand breaks. However, homologous recombination helps in first embryonic stages during the stages of cell replication [29,30].
Some studies have highlighted the role antioxidant therapy and varicocele repair to prevent sperm DNA fragmentation. Short abstinence periods and proper sperm handling in laboratories may prevent sperm DNA fragmentation. TESA –ICSI may also help to prevent sperm DNA fragmentation due to oxidative stress during transit in epididymis and vas deferens.
Sperm DNA defects can have several etiologies and mechanisms. Sperm DNA fragmentation analysis complements the routinely done semen analysis. The type, site and severity of sperm DNA defects as well as the oocyte repair capacity determine reproductive outcome. In future studies, various sperm selection criteria need to be compared. The mechanism of oocyte repair of sperm DNA defects also needs to be identified.
Sperm DNA fragmentation testing is a useful investigation in unexplained infertility. With an increase in paternal age the DNA fragmentation increases. Sperm count; motility and morphology have no significant relation with sperm DNA fragmentation index. In addition, Sperm DNA fragmentation has no significant association with Day 5 embryo grade in ICSI cycles. Future prospective studies with large sample size are required to confirm these preliminary findings.
The authors are thankful to ARC group of institutions and Saveetha Institute of Medical and Technical sciences for the providing the infrastructure and support for the conduct of study.
- Fleming S, Green S, Hall J. Analysis and alleviation of male infertility. Microsc Anal. 1995; 35: 37–39.
- Guzick DS, Overstreet JW, Factor-Litvak P, Brazil CK, Nakajima ST, et al. Sperm morphology, motility, and concentration in fertile and infertile men. N Engl J Med. 2001; 345: 1388–1393. PubMed: https://pubmed.ncbi.nlm.nih.gov/11794171/
- Agarwal A, Allamaneni SS. Sperm DNA damage assessment: a test whose time has come. Fertil Steril. 2005; 84: 850–853. PubMed: https://pubmed.ncbi.nlm.nih.gov/16213833/
- Irvine DS, Twigg JP, Gordon EL, Fulton N, Milne PA, et al. DNA integrity in human spermatozoa: relationships with semen quality. J Androl. 2000; 21: 33–44. PubMed: https://pubmed.ncbi.nlm.nih.gov/10670517/
- Agarwal A, Said TM. Role of sperm chromatin abnormalities and DNA damage in male infertility. Hum Reprod Update. 2003; 9: 331–345. PubMed: https://pubmed.ncbi.nlm.nih.gov/12926527/
- Zini A, Bielecki R, Phang D, Zenzes MT. Correlations between two markers of sperm DNA integrity, DNA denaturation and DNA fragmentation, in fertile and infertile men. Fertil Steril. 2001; 75: 674–677. PubMed:https://pubmed.ncbi.nlm.nih.gov/11287017/
- Oleszczuk K, Augustinsson L, Bayat N, Giwercman A, Bungum M. Prevalence of high DNA fragmentation index in male partners of unexplained infertile couples. Andrology. 2013; 1: 357–360. PubMed: https://pubmed.ncbi.nlm.nih.gov/23596042/
- Evgeni E, Lymberopoulos G, Gazouli M, Asimakopoulos B. Conventional semen parameters and DNA fragmentation in relation to fertility status in a Greek population. Eur J Obstet Gynecol Reprod Biol. 2015; 188: 17–23. PubMed: https://pubmed.ncbi.nlm.nih.gov/25770843/
- Zeqiraj A, Beadini S, Beadini N, Aliu H, Gashi Z, et al. Male Infertility and Sperm DNA Fragmentation. Open Access Maced J Med Sci. 2018; 6: 1342-1345. PubMed: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6108823/
- Diallo MS, Faye O, Diallo AS, Diallo Y, Diao B. Increased DNA fragmentation in patients with infertility in Dakar (Senegal) Advances in Reproductive Sciences. 2015; 3: 97.
- Yılmaz S, Zergeroğlu AD, Yılmaz E, Sofuoglu K, Delikara N, et al. Effects of sperm DNA fragmentation on semen parameters and ICSI outcome determined by an improved SCD test, Halosperm. Int J Fertil Steril. 2010; 4.
- Practice Committee of the American Society for Reproductive Medicine. Definitions of infertility and recurrent pregnancy loss: a committee opinion. Fertility and sterility. 2013; 99: 63. PubMed: https://pubmed.ncbi.nlm.nih.gov/23095139/
- Jungwirth A, Diemer T, Dohle GR, Giwercman A, Kopa Z, et al. Male Infertility. European Association of Urology. 2015; 62: 324–332. PubMed: https://pubmed.ncbi.nlm.nih.gov/22591628/
- O'Flynn O'Brien KL, Varghese AC, Agarwal A. The genetic causes of male factor infertility. Fertil Steril. 2010; 93: 1–12. PubMed: https://pubmed.ncbi.nlm.nih.gov/20103481/
- Twigg JP, Irvine DS, Aitken RJ. Oxidative damage to DNA in human spermatozoa does not preclude pronucleus formation at intracytoplasmic sperm injection. Hum Reprod. 1998; 13: 1864–1871. PubMed: https://pubmed.ncbi.nlm.nih.gov/9740440/
- Gorczyca W, Gong J, Darzynkiewicz Z. Detection of DNA strand breaks in individual apoptotic cells by the in situ terminal deoxynucleotidyl transferase and nick translation assays. Cancer Res. 1993; 53: 1945–1951. PubMed: https://pubmed.ncbi.nlm.nih.gov/8467513/
- Tarozzi N, Bizzaro D, Flamigni C, Borini A. Clinical relevance of sperm DNA damage in assisted reproduction. Reprod Biomed Online. 2007; 14: 746–757. PubMed: https://pubmed.ncbi.nlm.nih.gov/17579991/
- Sergerie M, Laforest G, Bujan L, Bissonnette F, Bleau G. Sperm DNA fragmentation: threshold value in male fertility. Hum Reprod. 2005;20: 3446–3451. PubMed: https://pubmed.ncbi.nlm.nih.gov/16085665/
- Oliva R. Protamines and male infertility. Hum Reprod Update. 2006; 12: 417-435. PubMed: https://pubmed.ncbi.nlm.nih.gov/16581810/
- Fernández-Díez C, González-Rojo S, Lombó M, Herráez MP. Impact of sperm DNA damage and oocyte-repairing capacity on trout development. Reproduction. 2016; 152: 57-67. PubMed: https://pubmed.ncbi.nlm.nih.gov/27071918/
- Pérez-Cerezales S, Martínez-Páramo S, Beirão J, Herráez MP. Fertilization capacity with rainbow trout DNA-damaged sperm and embryo developmental success. Reproduction. 2010; 139: 989-997. PubMed: https://pubmed.ncbi.nlm.nih.gov/20357047/
- Gunes S, Sertyel S. Sperm DNA Damage and Oocyte Repair Capability. A Clinician's Guide to Sperm DNA and Chromatin Damage, 2018.
- Zhao F, Yang Q, Shi S, Luo X, Sun Y. Semen Preparation Methods and Sperm Telomere Length: Density Gradient Centrifugation versus the Swim up Procedure. Sci. Rep. 2016; 6: 39051. PubMed: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5153621/
- Yang Q, Zhang N, Zhao F, Zhao W, Dai S, et al. Processing of Semen by Density Gradient Centrifugation Selects Spermatozoa with Longer Telomeres for Assisted Reproduction Techniques. Reprod BioMed Online. 2015; 31: 44–50. PubMed: https://pubmed.ncbi.nlm.nih.gov/25982091/
- Ménézo Y, Dale B, Cohen M. DNA damage and repair in human oocytes and embryos: a review. Zygote. 2010; 18: 357-365. PubMed: https://pubmed.ncbi.nlm.nih.gov/20663262/
- García-Rodríguez A, Gosálvez J, Agarwal A, Roy R, Johnston S. DNA Damage and Repair in Human Reproductive Cells. Int J Mol Sci. 2018; 20: 31. PubMed: https://pubmed.ncbi.nlm.nih.gov/30577615/
- Lord T, Aitken RJ. Fertilization Stimulates 8-Hydroxy-2′-Deoxyguanosine Repair and Antioxidant Activity to Prevent Mutagenesis in the Embryo. Dev. Biol. 2015; 406: 1–13. PubMed: https://pubmed.ncbi.nlm.nih.gov/26234752/
- Essers J, van Steeg H, de Wit J, Swagemakers SM, Vermeij M, et al. Homologous and non-homologous recombination differentially affect DNA damage repair in mice. EMBO J. 2000; 19: 1703–1710. PubMed: https://pubmed.ncbi.nlm.nih.gov/10747037/
- Rothkamm K, Krüger I, Thompson LH, Löbrich M. Pathways of DNA double-strand break repair during the mammalian cell cycle. Mol Cell Biol. 2003; 23: 5706–5715. PubMed: https://pubmed.ncbi.nlm.nih.gov/12897142/
- Derijck A, van der Heijden G, Giele M, Philippens M, de Boer P. DNA Double-Strand Break Repair in Parental Chromatin of Mouse Zygotes, the First Cell Cycle as an Origin of de novo Mutation. Hum. Mol. Genet. 2008; 17: 1922–1937. PubMed: https://pubmed.ncbi.nlm.nih.gov/18353795/